 
Amongst these mutants, the neutralization susceptibility of B.1.351 was significantly decreased to the immune sera from vaccinees ( 14– 16). In addition, several recent studies have focused on the neutralizing activity of vaccine-elicited humoral immunity against new circulating mutant lineages, including B.1.1.7 (United Kingdom, bearing mutations 69–70 del, 144 del, N501Y, A570D, D614G, P681H, T716I, S982A, and D1118H in the spike protein), B.1.429 (United States, bearing mutations S13I, W152C, L452R, and D614G in the spike protein), B.1.351(South Africa, bearing mutations D80A, D215G, K417N, E484K, N501Y D614G, and A701V in the spike protein), and P.1 and P.2 (Brazil, bearing certain mutations E484K, D614G, and V1176F, etc. However, a recent study indicated that the neutralizing activity of immune sera from vaccinees against G614 variant was better than the original one, and revealed that the structure of the G614 spike had a more open ACE2 binding site in the RBD region ( 11). This variant has a higher infectiousness due to its higher viral load in the respiratory secretions. Several studies demonstrated that the D614G mutation enhanced the replication of the mutated virus in the lung epithelial cells ( 10). 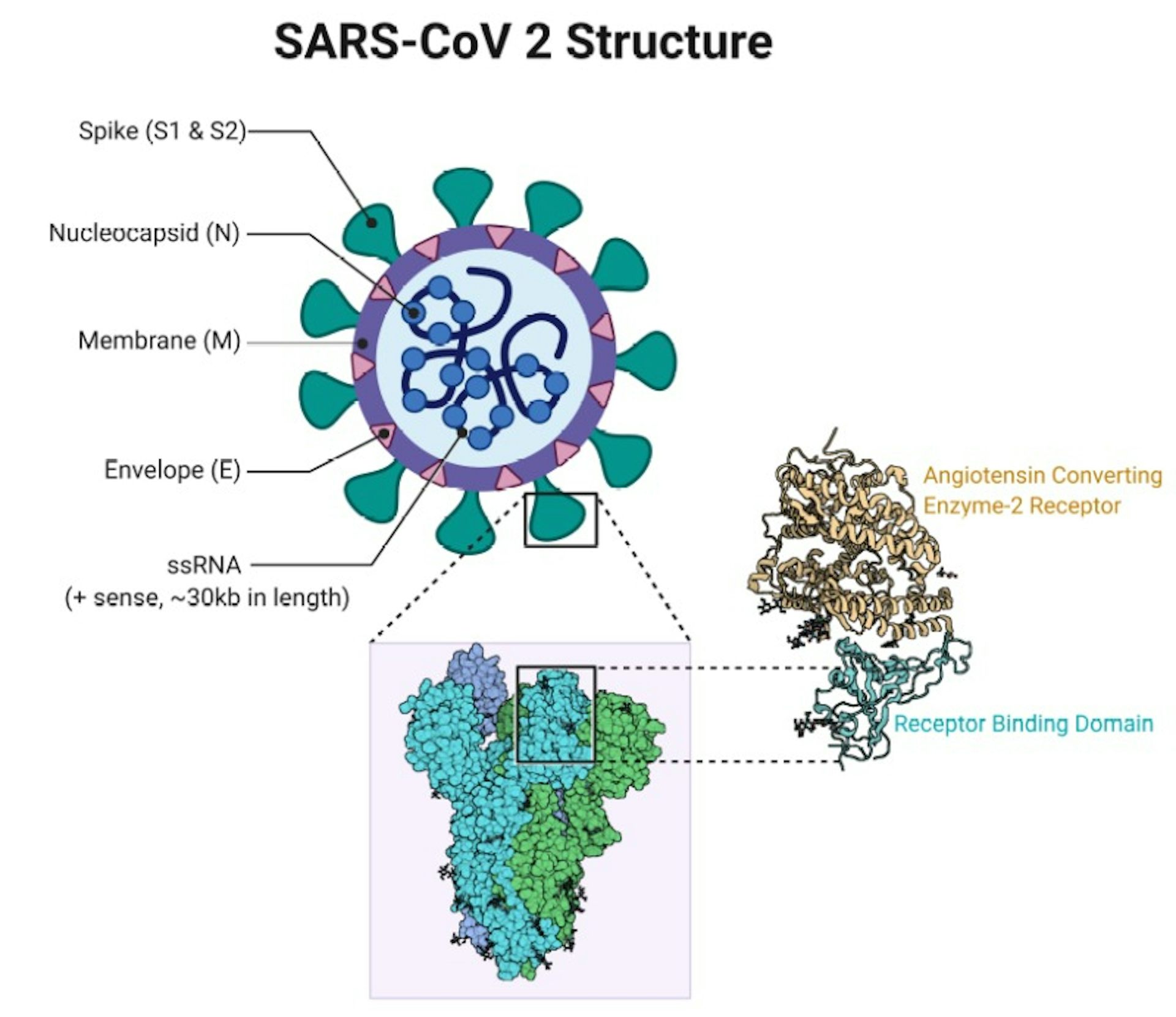
In contrast, early in the COVID-19 pandemic, SARS-CoV-2 spike mutation D614G rapidly became globally dominant. During the SARS epidemic in 2003, the single amino acid mutation D480A/G in the RBD domain of SARS-CoV spike protein gradually became dominant, and subsequent study confirmed that this mutation occurred in a critical site and enabled the mutant escaping neutralizing antibodies ( 9). 
The mutation in the spike protein not only affects the replication capacity and infectivity of the virus, but also impacts the host immune response. Importantly, the emergence of these amino acid mutations might result in a reduced neutralization ability of the immune serum from vaccinees or the existing antibodies in recovered COVID-19 patients, causing a new wave of the epidemic. Single amino acid mutation in the spike protein could affect the physicochemical property and structure of the protein, which subsequently alters the affinity of the receptor binding domain (RBD) with the viral receptor, human angiotensin-converting enzyme 2 (hACE2) ( 7, 8). However, the recent emergence of a new circulating mutant strain has raised public concern about the protection efficiency of the COVID-19 vaccine and antibody therapy. Therefore, the overall mutation frequency of SARS-CoV-2 genome is lower than that of HIV-1 and influenza viruses. Both of the two strategies have showed a remarkable effect on the control of the epidemic and/or the recovery of the patients ( 3, 4).ĭifferent from other RNA viruses, the non-structural protein nsp14 of SARS-CoV-2 has 3’-5’ exonuclease activity correcting the nucleotide mismatches occurred in the process of viral genome transcription ( 5, 6). Direct transfusion of specific SARS-CoV-2 antibodies, i.e., passive immunization, allows the individual to acquire immediate but short immunity, while the COVID-19 vaccine is able to elicit long-lasting immunity in the vaccinees. Since the coronavirus disease 2019 (COVID-19) pandemic is still spreading globally, specific antibody therapy and vaccine have now been approved to combat the causative agent SARS-CoV-2 ( 1, 2). 
A475V and E484Q mutants are highly resistant to neutralization by mAb B38 and 2-4, suggesting that some crucial mutations in spike protein might evolve SARS-CoV-2 variants capable of escaping humoral immune response. Three of 24 SARS-CoV-2 pseudoviruses containing different mutations in spike protein, including D614G, A475V, and E484Q, consistently showed an altered sensitivity to neutralization by convalescent plasmas. To identify crucial mutations in spike protein that escape humoral immune response, we evaluated the cross-neutralization of convalescent plasmas and RBD-specific monoclonal antibodies (mAbs) against various spike protein-based pseudoviruses. Small number of SARS-CoV-2 epidemic lineages did not efficiently exhibit a neutralization profile, while single amino acid mutation in the spike protein has not been confirmed in altering viral antigenicity resulting in immune escape. 2Department of Microbiology, Anhui Provincial Center for Disease Control and Prevention, Hefei, China.1The First Affiliated Hospital of USTC, Division of Life Sciences and Medicine, University of Science and Technology of China (USTC), Hefei, China.Chengchao Ding 1, Jun He 2, Xiangyu Zhang 1, Chengcheng Jiang 1, Yong Sun 2, Yuqing Zhang 1, Qingqing Chen 2, Hongliang He 1, Wenting Li 1, Jiajia Xie 1*, Zhirong Liu 2* and Yong Gao 1* 
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
